As we all know, a new crown pneumonia epidemic suddenly swept China at the end of 2019 and even affected the whole world. On January 30, 2020, WHO announced that the epidemic constituted a public health emergency of international concern (PHEIC).
The coronavirus discovered in Wuhan in December 2019 was named by the WHO as the 2019 Novel Coronavirus (2019-nCoV). On February 11, 2020, the virus was officially named SARS-CoV-2 by the Coronaviridae Study Group (CSG) of International Committee on Taxonomy of Viruses (ICTV). And the disease resulting from it is called COVID-19.
Overview of SARS-CoV-2
The newly discovered virus is 96% identical to bat coronavirus at the genome-wide level. Sequence analysis of paired proteins of seven conserved non-structural proteins showed that the virus belongs to the severe acute respiratory syndrome-related coronaviruses (SARSr-CoV), so it also belongs to the beta type of coronavirus family. The general incubation period of SARS-CoV-2 ranges from 2 to 14 days while the latency of SARS-CoV is usually about 3-7 days. The latest data shows that the longest incubation period reaches 24 days. After infected with SARS-CoV-2, the clinical symptoms of the patient are mainly fever, and there may be mild diarrhea, fatigue, and poor breathing. In addition, the virus is transmitted mainly through respiratory droplets and can be transmitted from person to person.
Invasion Mechanism of SARS-CoV-2
According to literature analysis, because SARS-CoV-2 and SARS-CoV are not only similar in their entire sequences, their S protein protein receptor-binding domain (RBD) sequences are also very similar. Therefore, it is likely that angiotensin-converting enzyme 2 (ACE2), the host receptor for SARS-CoV, is also a host receptor for SARS-CoV-2. Like the SARS-CoV virus, the S protein of SARS-CoV-2 first specifically binds to the receptor, and after the virus enters the cell, it begins to translate the RNA polymerase. It then transcribes its own genes into mRNAs and then translates other structural proteins. After translation, the virus enters the endoplasmic reticulum packaging, enters the Golgi apparatus for processing, and finally comes out of the cell by budding.
Treatment of SARS-CoV-2
Currently, there is no specific treatment for SARS-CoV-2, so the development of new drugs for this virus is extremely important. There are currently several possible strategies.
1. The first is the screening of existing broad-spectrum antiviral drugs. That is, screening of drugs that have been approved for the treatment of different viral infections to find drugs that can inhibit the SARS-CoV-2 life cycle as much as possible. However, this type of therapy cannot specifically treat SARS-CoV-2 and has certain side effects.
2. The second is to use existing molecular libraries and databases to screen for molecules that may have a therapeutic effect on this kind of coronavirus. This strategy can perform high-throughput screening in a short period of time and expand the scope of drug search. However, many drugs may show good anti-coronary virus activity in vitro, but have no value for human application in vivo, and the dosage and side effects of the drugs are unknown.
3. The third is to develop specific drugs based on the genomic information and pathological characteristics of SARS-CoV-2. This strategy can be divided into two categories, one acting on the human body and the other acting on the SARS-CoV-2.
- Action on the Human Body
It can be divided into three categories according to the location of action. In terms of the human immune system, the innate immune system response plays an important role in controlling the replication and infection of coronavirus. So, interferon is expected to achieve therapeutic purposes by enhancing the immune response. In human cells, blocking the signal pathways required for virus replication may play a certain antiviral effect. In order to enter host cells, viruses often bind to receptor proteins on the cell surface. Thus the development of drugs targeting these receptors can also have a good therapeutic effect.
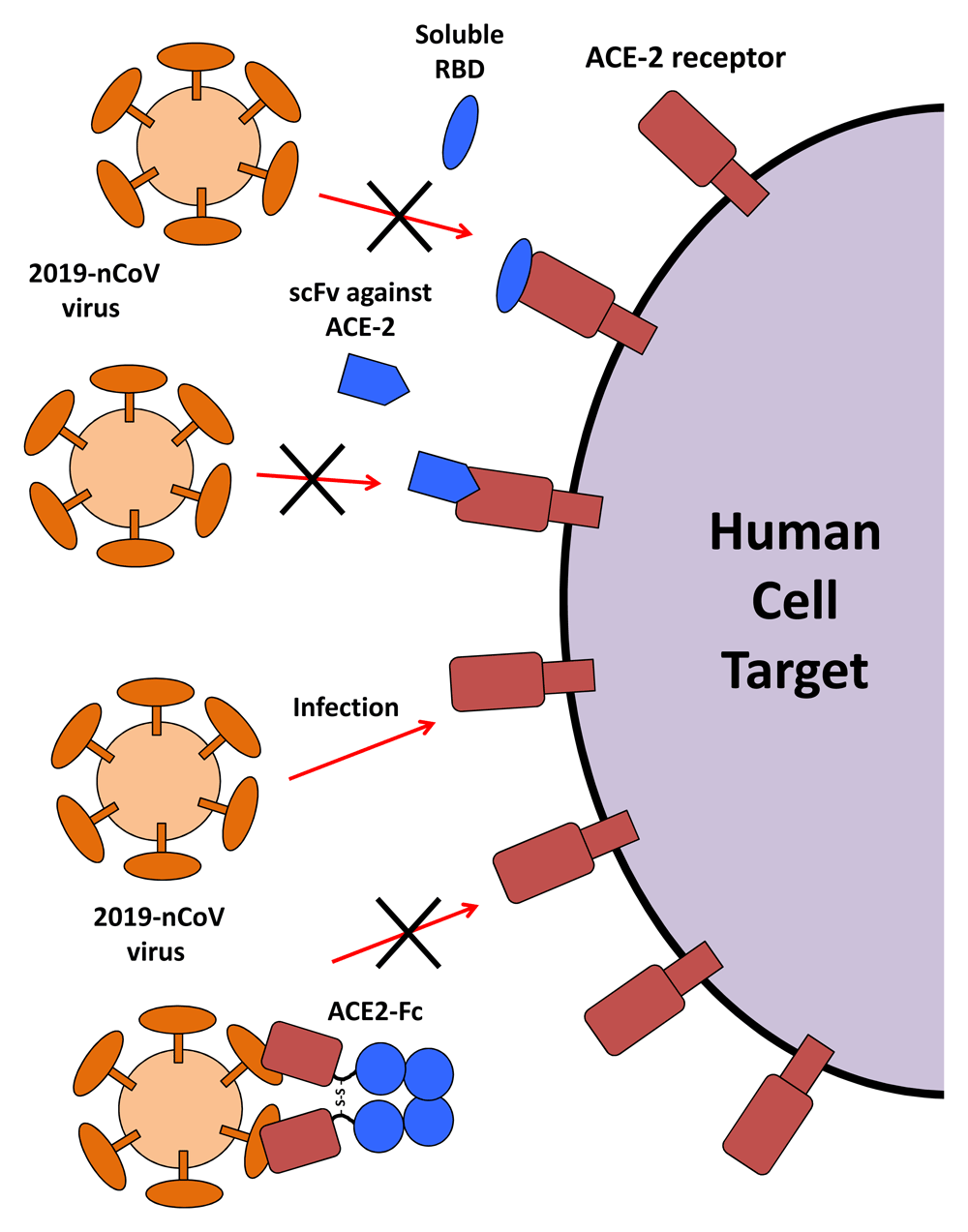
Because SARS-CoV-2 may bind to ACE2 on the host cell through the S protein and enter the cell, blocking the binding of S protein RBD to ACE2 can prevent the occurrence of infection. The possible treatments are shown in the figure below: the first one directly uses the RBD from the S protein of SARS-CoV or SARS-CoV-2 to bind ACE2 and occupy available sites. The second one directly uses ACE2 Antibodies or single-chain antibody fragments (scFv) to block the entry of the virus. The third type uses the RBD protein to fuse the Fc fragment to prevent the virus from infecting cells.
- Action on the SARS-CoV-2
Therapies that act on SARS-CoV-2 itself can be divided into several categories based on specific pathways of action. Act on the genetic material of the virus, thereby preventing viral RNA synthesis. Act on key enzymes of the virus, thereby further inhibiting virus replication. Act on the structural proteins of the virus, thereby blocking the virus’s binding to human cell receptors or inhibiting the virus’ self-assembly process.
Creative Biolabs is a leading service provider covering the entire antibody discovery and development process. We can provide you with coronavirus-related mouse/rabbit/human-derived recombinant antibodies, therapeutic antibodies and other related services to help your coronavirus drug development.
References:
[1] Wan Y, Shang J, Graham R, et al. Receptor recognition by novel coronavirus from Wuhan: An analysis based on decade-long structural studies of SARS[J]. Journal of virology, 2020.
[2] Xia S, Zhu Y, Liu M, et al. Fusion mechanism of 2019-nCoV and fusion inhibitors targeting HR1 domain in spike protein[J]. Cellular & Molecular Immunology, 2020: 1-3.
[3] Zumla A, Chan J F W, Azhar E I, et al. Coronaviruses—drug discovery and therapeutic options[J]. Nature reviews Drug discovery, 2016, 15(5): 327.
[4] Kruse R L. Therapeutic strategies in an outbreak scenario to treat the novel coronavirus originating in Wuhan, China[J]. F1000Research, 2020, 9(72): 72.