Cell-Free based Non-Canonical Amino Acid Antibody Labeling Service
The cell-free technique is a practical way to alter the target protein and give newly generated proteins more useful and extended functionalities. Target proteins can be co-translationally tagged with one or more non-canonical amino acids (ncAAs) by using cell-free protein synthesis. Cell-free systems offer an effective substitute because they are open systems unrestricted by the possible cytotoxicity of amino acid analogs, whereas in vivo methods are limited when a particular modified amino acid seems to have harmful side effects or when some amino acids simply cannot enter the cell. The naturally existing side chains of specific amino acids are used by artificial posttranslational modification technologies to perform chemical conjugation with a specific payload. Negative effects on protein solubility and functionality cannot be ruled out because the label is statistically incorporated into the protein at many places, producing a heterogeneous mixture of labeled proteins. Due to these limitations, methods for site-specific labeling have been developed, which make it easier to introduce a non-canonical amino acid at a single, predetermined point into a polypeptide chain.
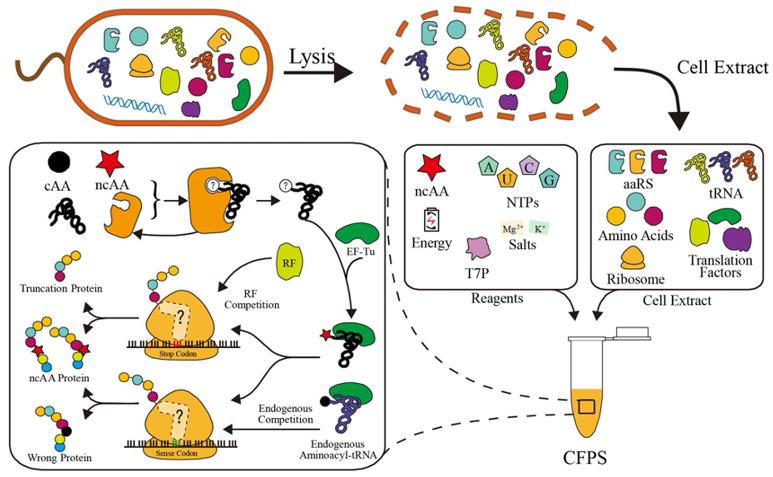 Fig.1. Schematic of cell-free system in ncAA incorporation.1
Fig.1. Schematic of cell-free system in ncAA incorporation.1
Labeling Strategies of Antibodies and Antibody Fragments with Non-canonical Amino Acids
- Genome Engineering
The most frequent reassignment for ncAA inclusion is the amber codon (UAG), which is typically a translation termination signal identified by peptide release factor 1 (RF1). The most straightforward method involves swapping out all of the amber codons for different stop codons to create cells that do not require RF1, allowing prfA to be deleted without impairing cell viability. One such E. Coli strain is C321. ΔA, in which all 321 of its amber codons have been swapped out for ochre codons (UAA). High-fidelity incorporation of ncAAs into proteins is possible in the absence of RF1 competition. For example, utilizing a cell-free system based on C321.ΔA cell extract, phosphoserine was effectively integrated into human MEK1 kinase, with robust output reaching up to milligram quantities.
- Protein Elimination
Protein elimination is a crucial strategy for labeling of antibodies and antibody fragments with ncAAs. Essential proteins can be kept active to promote cell growth by adjusting protein levels, or they can be eliminated to eliminate competition in the cell-free system. For instance, affinity tags, deactivating chemicals, or protease degradation can be used to eliminate RF1 selectively.
- Manipulation of tRNAs
A partially native protein is produced when a ncAA is inserted into a protein at a sense codon because the endogenous aminoacyl-tRNA will compete with the ncAA aminoacyl-tRNA in the ribosome during translation. While it is not practicable to directly edit tRNAs within the cell, cell-free system benefits from this approach since it removes the requirement for a membrane barrier, enabling tRNA modification to incorporate ncAA into a large number of sense codons. The easiest method is to replace each tRNA with a tRNA pool that has been specially made to meet the requirements of the study.
- Amino Acid Replacement
A few ncAAs that share structural similarities with canonical amino acids can be charged to native tRNAs by corresponding aaRS due to the relaxed amino acid specificity of aaRS. Selenomethionine, fluorinated tryptophan analogs, and chlorinated tyrosine analogs are a few examples. Therefore, the matching native amino acids can be replaced with structurally comparable non-coding amino acids (ncAAs). Without the need for designed aaRS or tRNAs, this amino acid replacement CFPS approach has proven to be an easy and affordable substitute for ncAA incorporation methodologies.
Creative Biolabs provides the world-leading cell-free antibody production service. In order to further the projects of our international clients, we additionally offer the highest quality labeling of antibodies and antibody fragments with non-canonical amino acids, based on our perfected process. Please feel free to contact us for more details.
Reference
- Wu, Yang, et al. "Emerging methods for efficient and extensive incorporation of non-canonical amino acids using cell-free systems." Frontiers in Bioengineering and Biotechnology 8 (2020): 863. Distributed under Open Access license CC BY 4.0, without modification.
For research use only. Not intended for any clinical use.
This site is protected by reCAPTCHA and the Google Privacy Policy and Terms of Service apply.

